Key Points
-
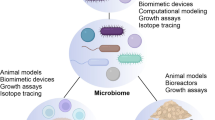
Whole metagenomic and metatranscriptomic sequencing endeavours are defining the functional potential and real-time activity of microbiomes and revealing interactions between microbial metabolism and host development.
-
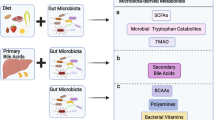
Gut microorganisms produce a diverse metabolite repertoire from the anaerobic fermentation of undigested dietary components that reach the colon, as well as from endogenous compounds that are generated by the microorganisms themselves and their hosts.
-
Complex carbohydrates are abundant substrates for bacterial fermentation in the colon, and their major metabolic end-products are short-chain fatty acids (SCFAs). SCFAs inhibit histone deacetylases (HDACs) and are ligands for G protein-coupled receptors; therefore, they act as signalling molecules that influence the expansion and function of haematopoietic and non-haematopoietic cell lineages. SCFA-driven HDAC inhibition tends to promote a tolerogenic, anti-inflammatory cell phenotype that is crucial for maintaining immune homeostasis and supports the concept that the microbiota can function as an epigenetic regulator of host physiology.
-
D-glycero-β-D-manno-heptose-1,7-bisphosphate (HBP), an intermediate in the lipopolysaccharide biosynthetic pathway of Gram-negative bacteria, initiates a novel innate immune signalling axis without first requiring bacterial lysis — a phenomenon that is so far unique to Neisseria gonorrhoeae.
-
Meta-omics and evolving computational frameworks are leading to the prediction and discovery of more microbial metabolites and components that are relevant to immune system function. It is also important to probe how well-known microbial metabolites (such as SCFAs) and co-metabolites (such as polyamines and aryl hydrocarbon receptor ligands) influence immune cell subsets and their functions.
Abstract
The microbiota — the collection of microorganisms that live within and on all mammals — provides crucial signals for the development and function of the immune system. Increased availability of technologies that profile microbial communities is facilitating the entry of many immunologists into the evolving field of host–microbiota studies. The microbial communities, their metabolites and components are not only necessary for immune homeostasis, they also influence the susceptibility of the host to many immune-mediated diseases and disorders. In this Review, we discuss technological and computational approaches for investigating the microbiome, as well as recent advances in our understanding of host immunity and microbial mutualism with a focus on specific microbial metabolites, bacterial components and the immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
We are sorry, but there is no personal subscription option available for your country.
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ding, T. & Schloss, P. D. Dynamics and associations of microbial community types across the human body. Nature 509, 357–360 (2014).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2015).
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Franzosa, E. A. et al. Sequencing and beyond: integrating molecular 'omics' for microbial community profiling. Nat. Rev. Microbiol. 13, 360–372 (2015).
Blekhman, R. et al. Host genetic variation impacts microbiome composition across human body sites. Genome Biol. 16, 191 (2015).
Morgan, X. C. & Huttenhower, C. Meta'omic analytic techniques for studying the intestinal microbiome. Gastroenterology 146, 1437–1448 (2014).
Dianda, L. et al. T cell receptor-αβ-deficient mice fail to develop colitis in the absence of a microbial environment. Am. J. Pathol. 150, 91–97 (1997).
Sellon, R. K. et al. Resident enteric bacteria are necessary for development of spontaneous colitis and immune system activation in interleukin-10- deficient mice. Infect. Immun. 66, 5224–5231 (1998).
Reháková, Z. et al. Germ-free mice do not develop ankylosing enthesopathy, a spontaneous joint disease. Hum. Immunol. 61, 555–558 (2000).
Sudo, N. et al. Postnatal microbial colonization programs the hypothalamic-pituitary-adrenal system for stress response in mice. J. Physiol. 558, 263–275 (2004).
Bohn, E. et al. Host gene expression in the colon of gnotobiotic interleukin-2-deficient mice colonized with commensal colitogenic or noncolitogenic bacterial strains: common patterns and bacteria strain specific signatures. Inflamm. Bowel Dis. 12, 853–862 (2006).
Wen, L. et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455, 1109–1113 (2008).
Wu, H.-J. et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32, 815–827 (2010).
Garrett, W. S. et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 8, 292–300 (2010).
Berer, K. et al. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature 479, 538–541 (2011).
Lee, Y. K., Menezes, J. S., Umesaki, Y. & Mazmanian, S. K. Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc. Natl Acad. Sci. USA 108 (Suppl. 1), 4615–4622 (2011).
Melkus, M. W. et al. Humanized mice mount specific adaptive and innate immune responses to EBV and TSST-1. Nat. Med. 12, 1316–1322 (2006).
Nochi, T., Denton, P. W., Wahl, A. & Garcia, J. V. Cryptopatches are essential for the development of human GALT. Cell Rep. 3, 1874–1884 (2013).
Laukens, D., Brinkman, B. M., Raes, J., De Vos, M. & Vandenabeele, P. Heterogeneity of the gut microbiome in mice: guidelines for optimizing experimental design. FEMS Microbiol. Rev. 40, 117–132 (2016).
Lichtman, J. S., Marcobal, A., Sonnenburg, J. L. & Elias, J. E. Host-centric proteomics of stool: a novel strategy focused on intestinal responses to the gut microbiota. Mol. Cell Proteom. 12, 3310–3318 (2013).
Marcobal, A. et al. A metabolomic view of how the human gut microbiota impacts the host metabolome using humanized and gnotobiotic mice. ISME J. 7, 1933–1943 (2013).
Sridharan, G. V. et al. Prediction and quantification of bioactive microbiota metabolites in the mouse gut. Nat. Commun. 5, 5492 (2014).
Kolmeder, C. A. & de Vos, W. M. Metaproteomics of our microbiome — developing insight in function and activity in man and model systems. J. Proteomics 97, 3–16 (2014).
Xiong, W., Abraham, P. E., Li, Z., Pan, C. & Hettich, R. L. Microbial metaproteomics for characterizing the range of metabolic functions and activities of human gut microbiota. Proteomics 15, 3424–3438 (2015).
Lasken, R. S. & McLean, J. S. Recent advances in genomic DNA sequencing of microbial species from single cells. Nat. Rev. Genet. 15, 577–584 (2014).
Cleary, B. et al. Detection of low-abundance bacterial strains in metagenomic datasets by eigengenome partitioning. Nat. Biotechnol. 33, 1053–1060 (2015).
Luo, C. et al. ConStrains identifies microbial strains in metagenomic datasets. Nat. Biotechnol. 33, 1045–1052 (2015).
Ufarté, L., Potocki-Veronese, G. & Laville, É. Discovery of new protein families and functions: new challenges in functional metagenomics for biotechnologies and microbial ecology. Front. Microbiol. 6, 563 (2015).
Earle, K. A. et al. Quantitative imaging of gut microbiota spatial organization. Cell Host Microbe 18, 478–488 (2015). This work showcases a robust pipeline for measuring the localization and organization of specific microorganisms and defined microbial communities within the gut.
Geva-Zatorsky, N. et al. In vivo imaging and tracking of host-microbiota interactions via metabolic labeling of gut anaerobic bacteria. Nat. Med. 21, 1091–1100 (2015). This paper describes an innovative strategy for labelling anaerobic microorganisms in vitro and tracking their location and function in vivo.
Hooper, L. V., Midtvedt, T. & Gordon, J. I. How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 22, 283–307 (2002).
Cummings, J. H., Pomare, E. W., Branch, W. J., Naylor, C. P. & Macfarlane, G. T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28, 1221–1227 (1987).
Usami, M. et al. Butyrate and trichostatin A attenuate nuclear factor κB activation and tumor necrosis factor α secretion and increase prostaglandin E2 secretion in human peripheral blood mononuclear cells. Nutr. Res. 28, 321–328 (2008).
Vinolo, M. A. R. et al. Suppressive effect of short- chain fatty acids on production of proinflammatory mediators by neutrophils. J. Nutr. Biochem. 22, 849–855 (2011).
Kendrick, S. F. W. et al. Acetate, the key modulator of inflammatory responses in acute alcoholic hepatitis. Hepatology 51, 1988–1997 (2010).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Singh, N. et al. Blockade of dendritic cell development by bacterial fermentation products butyrate and propionate through a transporter (Slc5a8)-dependent inhibition of histone deacetylases. J. Biol. Chem. 285, 27601–27608 (2010).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014). This study shows that gut microbiota-derived SCFAs from dietary fibre-enriched diets can dampen the severity of allergic inflammation in the lungs by signalling through GPR41 and generating tolerogenic DCs.
Tao, R. et al. Deacetylase inhibition promotes the generation and function of regulatory T cells. Nat. Med. 13, 1299–1307 (2007).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013). This paper establishes that enhanced HDAC activity at the Foxp3 locus is one mechanism for how microbial-derived butyrate induces the differentiation and suppressive function of peripheral T reg cells and how colonic T reg cells can protect against the development of colitis.
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013). This work demonstrates that butyrate specifically promotes the differentiation of peripheral, not thymus-derived, T reg cells and that the mode of delivery, local versus systemic administration, influences SCFA-mediated effects on colonic T reg cells.
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013). This paper illustrates that microbiota-derived SCFAs regulate T reg cell homeostasis in the colon and that SCFA-mediated effects on T reg cells are mediated in part by GPR43.
Thorburn, A. N. et al. Evidence that asthma is a developmental origin disease influenced by maternal diet and bacterial metabolites. Nat. Commun. 6, 7320 (2015).
Maslowski, K. M. & Mackay, C. R. Diet, gut microbiota and immune responses. Nat. Immunol. 12, 5–9 (2011).
Wang, L., de Zoeten, E. F., Greene, M. I. & Hancock, W. W. Immunomodulatory effects of deacetylase inhibitors: therapeutic _targeting of FOXP3+ regulatory T cells. Nat. Rev. Drug Discov. 8, 969–981 (2009).
Thorburn, A. N., Macia, L. & Mackay, C. R. Diet, metabolites, and 'western-lifestyle' inflammatory diseases. Immunity 40, 833–842 (2014).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009). This landmark paper provides a molecular link between diet, the gut microbiota and host immune function by establishing that GPR43 expression on immune cells is crucial for imparting the regulatory properties of SCFAs and for resolving inflammation in experimental models of colitis, arthritis and asthma.
Voltolini, C. et al. A novel antiinflammatory role for the short-chain fatty acids in human labor. Endocrinology 153, 395–403 (2012).
Vieira, A. T. et al. A role for gut microbiota and the metabolite-sensing receptor GPR43 in a murine model of gout. Arthritis Rheumatol 67, 1646–1656 (2015).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015). This paper demonstrates that microbiota-generated SCFAs from the gut regulate microglia homeostasis, which has important implications for local and systemic consequences of microbial dysbiosis on CNS function.
Singh, N. et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 40, 128–139 (2014). This study illustrates that the SCFA receptor GPR109A on immune and epithelial cells triggers the production of protective cytokines and promotes anti-inflammatory immune responses, which are crucial for preventing colitis and colitis-associated colorectal cancer.
Ghorbani, P. et al. Short-chain fatty acids affect cystic fibrosis airway inflammation and bacterial growth. Eur. Respir. J. 46, 1033–1045 (2015).
Willemsen, L. E. M., Koetsier, M. A., van Deventer, S. J. H. & van Tol, E. A. F. Short chain fatty acids stimulate epithelial mucin 2 expression through differential effects on prostaglandin E1 and E2 production by intestinal myofibroblasts. Gut 52, 1442–1447 (2003).
Gaudier, E. et al. Butyrate specifically modulates MUC gene expression in intestinal epithelial goblet cells deprived of glucose. Am. J. Physiol. Gastrointest. Liver Physiol. 287, G1168–G1174 (2004).
Wrzosek, L. et al. Bacteroides thetaiotaomicron and Faecalibacterium prausnitzii influence the production of mucus glycans and the development of goblet cells in the colonic epithelium of a gnotobiotic model rodent. BMC Biol. 11, 61 (2013).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011).
Macia, L. et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 6, 6734 (2015).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Schilderink, R., Verseijden, C. & de Jonge, W. J. Dietary inhibitors of histone deacetylases in intestinal immunity and homeostasis. Front. Immunol. 4, 226 (2013).
Wu, J., Zhou, Z., Hu, Y. & Dong, S. Butyrate-induced GPR41 activation inhibits histone acetylation and cell growth. J. Genet. Genom. 39, 375–384 (2012).
Li, Y. et al. Exogenous stimuli maintain intraepithelial lymphocytes via aryl hydrocarbon receptor activation. Cell 147, 629–640 (2011).
Kiss, E. A. et al. Natural aryl hydrocarbon receptor ligands control organogenesis of intestinal lymphoid follicles. Science 334, 1561–1565 (2011).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013). This paper offers a mechanistic insight into the functional role of microbial-derived tryptophan metabolites in gut immune homeostasis through activation of the AHR in ILC3s, which provides colonization resistance to a fungal pathogen and protection from mucosal inflammation.
Hubbard, T. D. et al. Adaptation of the human aryl hydrocarbon receptor to sense microbiota-derived indoles. Sci. Rep. 5, 12689 (2015).
Di Martino, M. L. et al. Polyamines: emerging players in bacteria-host interactions. Int. J. Med. Microbiol. 303, 484–491 (2013).
Chen, J. et al. Polyamines are required for expression of Toll-like receptor 2 modulating intestinal epithelial barrier integrity. Am. J. Physiol. Gastrointest. Liver Physiol. 293, G568–G576 (2007).
Liu, L. et al. Polyamines regulate E-cadherin transcription through c-Myc modulating intestinal epithelial barrier function. Am. J. Physiol. Cell Physiol. 296, C801–C810 (2009).
Dufour, C. et al. Spermine and spermidine induce intestinal maturation in the rat. Gastroenterology 95, 112–116 (1988).
Buts, J. P., De Keyser, N., Kolanowski, J., Sokal, E. & Van Hoof, F. Maturation of villus and crypt cell functions in rat small intestine. Role of dietary polyamines. Dig. Dis. Sci. 38, 1091–1098 (1993).
Löser, C., Eisel, A., Harms, D. & Fölsch, U. R. Dietary polyamines are essential luminal growth factors for small intestinal and colonic mucosal growth and development. Gut 44, 12–16 (1999).
Zhang, M., Wang, H. & Tracey, K. J. Regulation of macrophage activation and inflammation by spermine: a new chapter in an old story. Crit. Care Med. 28, N60–N66 (2000).
Kibe, R. et al. Upregulation of colonic luminal polyamines produced by intestinal microbiota delays senescence in mice. Sci. Rep. 4, 4548 (2014).
Perez-Cano, F. J., González-Castro, A., Castellote, C., Franch, A. & Castell, M. Influence of breast milk polyamines on suckling rat immune system maturation. Dev. Comp. Immunol. 34, 210–218 (2010).
Minois, N., Carmona-Gutierrez, D. & Madeo, F. Polyamines in aging and disease. Aging (Albany NY) 3, 716–732 (2011).
Matsumoto, M., Kurihara, S., Kibe, R., Ashida, H. & Benno, Y. Longevity in mice is promoted by probiotic-induced suppression of colonic senescence dependent on upregulation of gut bacterial polyamine production. PLoS ONE 6, e23652 (2011).
Miller-Fleming, L., Olin-Sandoval, V., Campbell, K. & Ralser, M. Remaining mysteries of molecular biology: the role of polyamines in the cell. J. Mol. Biol. 427, 3389–3406 (2015).
Gerner, E. W. & Meyskens, F. L. Polyamines and cancer: old molecules, new understanding. Nat. Rev. Cancer 4, 781–792 (2004).
Johnson, C. H. et al. Metabolism links bacterial biofilms and colon carcinogenesis. Cell. Metab. 21, 891–897 (2015). This work demonstrates a role for colonic mucosal biofilms in potentiating colonic carcinogenesis through the generation of polyamine metabolites that can enhance cell proliferation and cancer cell growth.
Hayes, C. S. et al. Polyamine-blocking therapy reverses immunosuppression in the tumor microenvironment. Cancer Immunol. Res. 2, 274–285 (2014).
Wang, Q. et al. A bacterial carbohydrate links innate and adaptive responses through Toll-like receptor 2. J. Exp. Med. 203, 2853–2863 (2006).
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Dasgupta, S., Erturk-Hasdemir, D., Ochoa-Repáraz, J., Reinecker, H.-C. & Kasper, D. L. Plasmacytoid dendritic cells mediate anti-inflammatory responses to a gut commensal molecule via both innate and adaptive mechanisms. Cell Host Microbe 15, 413–423 (2014).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Round, J. L. et al. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science 332, 974–977 (2011).
Wang, Y. et al. A commensal bacterial product elicits and modulates migratory capacity of CD39+ CD4 T regulatory subsets in the suppression of neuroinflammation. Gut Microbes 5, 552–561 (2014).
Dwyer, K. M. et al. CD39 and control of cellular immune responses. Purinerg. Signal. 3, 171–180 (2007).
Telesford, K. M. et al. A commensal symbiotic factor derived from Bacteroides fragilis promotes human CD39+Foxp3+ T cells and Treg function. Gut Microbes 6, 234–242 (2015).
Gibson, D. J. et al. Heightened expression of CD39 by regulatory T lymphocytes is associated with therapeutic remission in inflammatory bowel disease. Inflamm. Bowel Dis. 21, 2806–2814 (2015).
Liu, M. et al. Formylpeptide receptors mediate rapid neutrophil mobilization to accelerate wound healing. PLoS ONE 9, e90613 (2014).
Kretschmer, D., Rautenberg, M., Linke, D. & Peschel, A. Peptide length and folding state govern the capacity of staphylococcal β-type phenol-soluble modulins to activate human formyl-peptide receptors 1 or 2. J. Leukoc. Biol. 97, 689–697 (2015).
Bloes, D. A., Kretschmer, D. & Peschel, A. Enemy attraction: bacterial agonists for leukocyte chemotaxis receptors. Nat. Rev. Microbiol. 13, 95–104 (2015).
Chiu, I. M. et al. Bacteria activate sensory neurons that modulate pain and inflammation. Nature 501, 52–57 (2013). This paper illuminates an unanticipated role for the CNS in mediating host–pathogen interactions, by showing that a bacterial pathogen can stimulate pain and influence inflammatory immune responses by directly activating peripheral sensory neurons.
Haas, P.-J. et al. N-terminal residues of the chemotaxis inhibitory protein of Staphylococcus aureus are essential for blocking formylated peptide receptor but not C5a receptor. J. Immunol. 173, 5704–5711 (2004).
Prat, C. et al. A homolog of formyl peptide receptor-like 1 (FPRL1) inhibitor from Staphylococcus aureus (FPRL1 inhibitory protein) that inhibits FPRL1 and FPR. J. Immunol. 183, 6569–6578 (2009).
Yang, S. et al. Synthesis, antibacterial activity, and biological evaluation of formyl hydroxyamino derivatives as novel potent peptide deformylase inhibitors against drug-resistant bacteria. Eur. J. Med. Chem. 86, 133–152 (2014).
Lewandowski, T. et al. Staphylococcus aureus formyl-methionyl transferase mutants demonstrate reduced virulence factor production and pathogenicity. Antimicrob. Agents Chemother. 57, 2929–2936 (2013).
Min, S. et al. Frequency of spontaneous resistance to peptide deformylase inhibitor GSK1322322 in Haemophilus influenzae, Staphylococcus aureus, Streptococcus pyogenes, and Streptococcus pneumoniae. Antimicrob. Agents Chemother. 59, 4644–4652 (2015).
Gaudet, R. G. et al. Cytosolic detection of the bacterial metabolite HBP activates TIFA-dependent innate immunity. Science 348, 1251–1255 (2015). This work identifies a novel innate immune signalling axis that is initiated by a pro-inflammatory bacterial-derived metabolite, an intermediate in lipopolysaccharide biosynthesis of Gram-negative bacteria.
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS ONE 5, e14004 (2010).
Burbano, H. A. et al. _targeted investigation of the Neandertal genome by array-based sequence capture. Science 328, 723–725 (2010).
Duncavage, E. J. et al. Hybrid capture and next-generation sequencing identify viral integration sites from formalin-fixed, paraffin-embedded tissue. J. Mol. Diagn. 13, 325–333 (2011).
Cabanski, C. R. et al. cDNA hybrid capture improves transcriptome analysis on low-input and archived samples. J. Mol. Diagn. 16, 440–451 (2014).
Lim, S. W., Tran, T. M. & Abate, A. R. PCR-activated cell sorting for cultivation-free enrichment and sequencing of rare microbes. PLoS ONE 10, e0113549 (2015).
Maurice, C. F. & Turnbaugh, P. J. Quantifying the metabolic activities of human-associated microbial communities across multiple ecological scales. FEMS Microbiol. Rev. 37, 830–848 (2013).
Segata, N. et al. Computational meta'omics for microbial community studies. Mol. Syst. Biol. 9, 666 (2013).
Faith, J. J. et al. Creating and characterizing communities of human gut microbes in gnotobiotic mice. ISME J. 4, 1094–1098 (2010).
Wostmann, B. S., Larkin, C., Moriarty, A. & Bruckner-Kardoss, E. Dietary intake, energy metabolism, and excretory losses of adult male germfree Wistar rats. Lab. Anim. Sci. 33, 46–50 (1983).
Rosenbaum, M., Knight, R. & Leibel, R. L. The gut microbiota in human energy homeostasis and obesity. Trends Endocrinol. Metab. 26, 493–501 (2015).
Deplancke, B. & Gaskins, H. R. Microbial modulation of innate defense: goblet cells and the intestinal mucus layer. Am. J. Clin. Nutr. 73, 1131S–1141S (2001).
Smith, K., McCoy, K. D. & Macpherson, A. J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 19, 59–69 (2007).
Pabst, O. New concepts in the generation and functions of IgA. Nat. Rev. Immunol. 12, 821–832 (2012).
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Strauch, U. G. et al. Influence of intestinal bacteria on induction of regulatory T cells: lessons from a transfer model of colitis. Gut 54, 1546–1552 (2005).
Belkaid, Y. & Naik, S. Compartmentalized and systemic control of tissue immunity by commensals. Nat. Immunol. 14, 646–653 (2013).
Borre, Y. E. et al. Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol. Med. 20, 509–518 (2014).
Belcheva, A. et al. Gut microbial metabolism drives transformation of MSH2-deficient colon epithelial cells. Cell 158, 288–299 (2014).
Donohoe, D. R. et al. A gnotobiotic mouse model demonstrates that dietary fiber protects against colorectal tumorigenesis in a microbiota- and butyrate-dependent manner. Cancer Discov. 4, 1387–1397 (2014).
Srinivasan, S. et al. Metabolic signatures of bacterial vaginosis. MBio 6, e00204–00215 (2015).
Koeth, R. A. et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013).
Pluznick, J. L. et al. Olfactory receptor responding to gut microbiota-derived signals plays a role in renin secretion and blood pressure regulation. Proc. Natl Acad. Sci. USA 110, 4410–4415 (2013).
Meijers, B. K. I. et al. p-Cresol and cardiovascular risk in mild-to-moderate kidney disease. Clin. J. Am. Soc. Nephrol. 5, 1182–1189 (2010).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Braniste, V. et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci. Transl Med. 6, 263ra158 (2014).
Buffie, C. G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015).
Acknowledgements
The authors thank the members of the Garrett laboratory for their helpful discussions. The work relevant to this Review is supported by the grants R01 CA154426 and R01 GM099531, a Burroughs Wellcome Career in Medical Sciences Award and a Searle Scholars Award to W.S.G.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Dysbiosis
-
An imbalance in the composition or function of the microbial species that are normally found in mammalian hosts. It is associated with alterations in immune function and susceptibility to inflammatory diseases, allergies and metabolic conditions.
- Gnotobiotic mouse models
-
Experimental models in which germ-free mice are selectively colonized by defined microorganisms and kept in isolators to control their microbial colonization status.
- Bone marrow–liver–thymus humanized mice
-
(BLT humanized mice). Immunodeficient mice that are engrafted with human fetal liver and thymus under the renal capsule. Three weeks later, mice are irradiated and then injected with a suspension of CD34+ cells from the same human fetal liver sample. These fetal liver cells seed to the mouse bone marrow.
- Humanized mouse models
-
Experimental models in which mice carry functioning human genes, cells, tissues (including faecal material) or organs that are introduced by transgenesis, injection or transplantation. For example, an immunodeficient mouse transgenically expressing susceptibility genes for type 1 diabetes and reconstituted with T cells from a patient with type 1 diabetes and human islets of Langerhans can be used to study relevant autoimmune processes.
- Fluorescence in situ hybridization
-
(FISH). A technique in which fluorescent probes are used to visually label specific DNA sequences in the nuclei of cells.
- Metabolic oligosaccharide engineering
-
(MOE). A technology in which synthetic sugar analogues are exogenously supplied to living cells and biosynthetically incorporated into cell surface polysaccharides. An advantage of this technology is that it can be used for prokaryotic and eukaryotic cells that are grown under aerobic or anaerobic conditions.
- Bio-orthogonal click chemistry
-
(BCC). A chemical reaction in a living cell that allows a labelled synthetic probe to be covalently linked to _targeted cellular substrates without disrupting any native functions of the cell. This method can be used to tag and visualize biomolecules within cells of interest.
- Anaerobic fermentation
-
The process of extracting energy from carbohydrates. Some bacteria are facultative anaerobes, meaning they can switch between aerobic respiration and anaerobic pathways, depending on the availability of oxygen or other electron acceptors. Other bacteria are obligate anaerobes, meaning they completely rely on anaerobic fermentation and can only survive in the absence of oxygen.
- Histone deacetylases
-
(HDACs). Enzymes that remove the acetyl groups from lysine residues that are located at the amino termini of histones. In general, decreased levels of histone acetylation are associated with the repression of gene expression. The balance of histone acetylation is maintained by the interplay between HDACs and histone acetyltransferases.
- Dextran sodium sulfate-induced injury model
-
(DSS-induced injury model). A commonly used experimental model of colonic injury and mucosal inflammation induced in mice by ingestion of the sulfated polysaccharide DSS. This model causes acute colonic epithelial damage and inflammation.
- T cell transfer colitis model
-
A well-characterized model of chronic colitis that is induced by the transfer of CD4+CD45RBhi (naive) T cells from healthy wild-type mice into immunodeficient syngeneic recipients.
- Xenobiotics
-
Compounds inclusive of drugs, food components and pollutants.
- Innate lymphoid cell
-
(ILC). A lymphoid cell that is derived from the common lymphoid progenitor and does not express a recombined antigen receptor. ILCs have important roles in innate immune responses to infectious microorganisms, in epithelial homeostasis and in lymphoid tissue formation.
Rights and permissions
About this article
Cite this article
Rooks, M., Garrett, W. Gut microbiota, metabolites and host immunity. Nat Rev Immunol 16, 341–352 (2016). https://doi.org/10.1038/nri.2016.42
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri.2016.42