Comparative Analysis for Genetic Characterization in Korean Native Jeju Horse
- PMID: 34203473
- PMCID: PMC8300358
- DOI: 10.3390/ani11071924
Comparative Analysis for Genetic Characterization in Korean Native Jeju Horse
Abstract
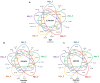
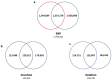
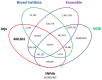
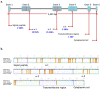
The Jeju horse is a native Korean species that has been breeding on Jeju Island since the 13th century. Their shape has a distinct appearance from the representative species, Thoroughbred. Here, we performed a comparison of the Jeju horse and Thoroughbred horse for the identification of genome-wide structure variation by using the next-generation sequencing (NGS) technique. We generated an average of 95.59 Gb of the DNA sequence, resulting in an average of 33.74 X sequence coverage from five Jeju horses. In addition, reads obtained from WGRS data almost covered the horse reference genome (mapped reads 98.4%). Based on our results, we identified 1,244,064 single nucleotide polymorphisms (SNPs), 113,498 genomic insertions, and 114,751 deletions through bioinformatics analysis. Interestingly, the results of the WGRS comparison indicated that the eqCD1a6 gene contains signatures of positive natural selection in Jeju horses. The eqCD1a6 gene is known to be involved in immunity. The eqCD1a6 gene of Jeju horses commonly contained 296 variants (275 SNPs and 21 INDELs) that were compared with its counterpart of two Thoroughbred horses. In addition, we used LOAA, digital PCR, to confirm the possibility of developing a molecular marker for species identification using variant sites. As a result, it was possible to confirm the result of the molecular marker with high accuracy. Nevertheless, eqCD1a6 was shown to be functionally intact. Taken together, we have found significant genomic variation in these two different horse species.
Keywords: Equus ferus caballus; Jeju horse; SNP; Thoroughbred; WGRS; digital PCR; eqCD1a6.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


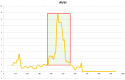
Similar articles
-
Genome-wide analyses of the Jeju, Thoroughbred, and Jeju crossbred horse populations using the high density SNP array.Genes Genomics. 2018 Nov;40(11):1249-1258. doi: 10.1007/s13258-018-0722-0. Epub 2018 Aug 11. Genes Genomics. 2018. PMID: 30099720
-
Comprehensive genome and transcriptome analyses reveal genetic relationship, selection signature, and transcriptome landscape of small-sized Korean native Jeju horse.Sci Rep. 2019 Nov 13;9(1):16672. doi: 10.1038/s41598-019-53102-8. Sci Rep. 2019. PMID: 31723199 Free PMC article.
-
Whole genome sequencing analysis of horse populations inhabiting the Korean Peninsula and Przewalski's horse.Genes Genomics. 2019 Jun;41(6):621-628. doi: 10.1007/s13258-019-00795-w. Epub 2019 Apr 2. Genes Genomics. 2019. PMID: 30941726
-
The genetics of equine osteochondrosis.Vet J. 2013 Jul;197(1):13-8. doi: 10.1016/j.tvjl.2013.03.036. Epub 2013 Jul 1. Vet J. 2013. PMID: 23809989 Review.
-
Ten years of the horse reference genome: insights into equine biology, domestication and population dynamics in the post-genome era.Anim Genet. 2019 Dec;50(6):569-597. doi: 10.1111/age.12857. Epub 2019 Sep 30. Anim Genet. 2019. PMID: 31568563 Free PMC article. Review.
References
-
- Petersen J.L., Mickelson J.R., Rendahl A.K., Valberg S.J., Andersson L.S., Axelsson J., Bailey E., Bannasch D., Binns M.M., Borges A.S., et al. Genome-wide analysis reveals selection for important traits in domestic horse breeds. PLoS Genet. 2013;9:e1003211. doi: 10.1371/journal.pgen.1003211. - DOI - PMC - PubMed
-
- Zhou M., Wang Q., Sun J., Li X., Xu L., Yang H., Shi H., Ning S., Chen L., Li Y., et al. In silico detection and characteristics of novel microRNA genes in the Equus caballus genome using an integrated ab initio and comparative genomic approach. Genomics. 2009;94:125–131. doi: 10.1016/j.ygeno.2009.04.006. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

